Article Content
Abstract
Background:
Drug recommendation is a crucial application of artificial intelligence in medical practice. Although many models have been proposed to solve this task, two challenges remain unresolved: (i) most existing models use all historical visits as input, overlooking fine-grained correlations between historical and current information; (ii) Electronic Health Records (EHRs) are underutilized, with only partial information considered to describe patient conditions. To tackle the challenges, we propose a novel drug recommendation model, denoted by EDRMM, which incorporates multi-granularity and multi-attribute information into representation learning. We develop a longitudinal attribute-level history selection mechanism to effectively identify fine-grained historical information that is highly relevant to a patient’s current clinical conditions. We analyze the impact of key Electronic Health Record (EHR) attributes, demonstrating that incorporating such attributes into patient representations can further boost performance. We also design an adaptive global Drug–Drug Interaction (DDI) risk regularization term for the DDI loss function to better balance accuracy and safety during training.
Results:
Experimental results show that our model achieves state-of-the-art performance on a widely used MIMIC-III dataset.
Conclusions:
EDRMM overcomes two key drug recommendation limitations through three innovations: (1) Dynamic attribute-level history selection, which retrieves relevant features and filters out noise, (2) The integration of multi-attribute EHR with attribute-specific encoding strategies to generate comprehensive patient representations, and (3) Hybrid optimization balancing accuracy and safety via adaptive DDI regularization. The combination of these three innovations enables the proposed EDRMM to achieve the best recommendation performance on the MIMIC-III dataset.
Background
Introduction
Issues such as adverse drug reactions, drug interactions, and incomplete patient medical histories highlight the critical need for advanced decision support. As shown in Fig. 1, the input to the drug recommendation task comes from EHRs. By leveraging artificial intelligence, the model can provide accurate, efficient, and personalized drug prescriptions[1,2,3]. Recently, deep learning has shown its efficacy in drug recommendation tasks [4,5,6,7,8,9,10,11]. Existing work can be categorized into two groups based on the temporal scope of the data used. The first category is instance-based models, which recommend drugs based on the patient’s current condition without considering historical information [12, 13]. However, they ignore the historical information that may be useful for modeling the patient’s current clinical conditions. The longitudinal-based models integrate both the patient’s historical and current conditions, capturing long-term states [4, 5, 8].
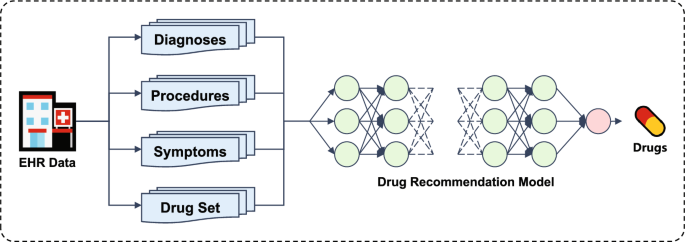
The framework of drug recommendation task
Although moderate progress has been made, two challenges remain unresolved. (i) The relationship between historical and current visits is ignored. Existing models typically use all historical data to learn global patient representations. However, for patients with complex and chronic conditions, only certain visit records may be relevant to their current clinical state. The use of global representations can overlook these important associations, introducing irrelevant information and reducing model performance. Therefore, capturing the fine-grained correlations between historical and current information is a key challenge in improving drug recommendation performance. (ii) EHRs are underutilized, with only partial information considered to describe patient conditions. EHRs typically contain a wide range of attributes, such as diagnoses, procedures, and symptoms. However, most existing models focus on leveraging historical diagnosis and procedure information for patient representation. Most models rely on only one or two attributes, and they fail to utilize the rich information available in EHRs fully (seen in Table 1). Although DrugRec [11] incorporates three attributes, it has not explored the detailed impacts of each attribute.
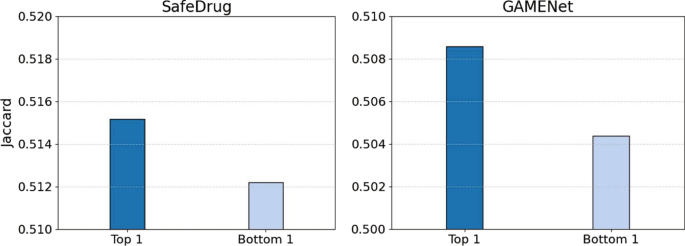
Experimental results for SafeDrug (left) and GAMENet (right)
In Fig. 2, we conducted experiments on two models of GAMENet [4] and SafeDrug [5] in the MIMIC-III dataset, by using the Jaccard similarity metric for evaluation. For the multi-visit data, we first quantify the relevance by calculating the Jaccard similarity of diagnoses between two EHRs. We then evaluate the model performance by inputting either the most relevant (Top 1) or the most irrelevant (Bottom 1) historical data. The results indicate that models using the most relevant historical information (Top 1) significantly outperform those using the most irrelevant information (Bottom 1). This finding points out the critical role of historical information in enhancing the effectiveness of drug recommendation models.
To address the aforementioned challenges, we propose EDRMM, which is a novel drug recommendation model that employs multi-granularity and multi-attribute historical information for EHR representation learning. In addition, to balance the accuracy and safety of drug recommendations, we design a global DDI risk regularization term for DDI loss, which incorporates global DDI risk knowledge to capture potential drug interaction patterns. Our main contributions are as follows:
- To capture the correlation between historical and current visits, we develop an attribute-level history selection mechanism to retrieve relevant information while filtering out noise.
- To fully utilize the significant information from EHRs, we explore the effects of different attributes (i.e., diagnosis, procedure, and symptom) on the performance of drug recommendation.
- We design a global DDI risk regularization term for the DDI loss function, which enhances the model by effectively capturing global drug interaction patterns and mitigating potential DDI risks.
- We conduct extensive experiments on the MIMIC-III dataset to compare EDRMM with several strong baseline models. The results demonstrate the superiority of EDRMM.
Related work
Existing drug recommendation models can be categorized into two groups: instance-based and longitudinal-based models.
Instance-based models
RETAIN [14] emphasizes model interpretability through a two-level attention mechanism to capture patient behavior. LEAP [13] treats drug recommendation as a sequential decision-making problem, utilizing recurrent decoders to model label dependencies.
Longitudinal-based methods
GAMENet [4] introduces graph-enhanced memory networks that integrate multiple knowledge graphs using a late-fusion mechanism. SafeDrug [5] combines a global message-passing neural network module with a local bipartite learning module to comprehensively encode drug molecular connectivity and functionality. MICRON [6] focuses on identifying drug changes using a recurrent architecture. DrugRec [11] employs a causal graphical model with front-door adjustment to mitigate hidden recommendation biases. COGNet [9] implements a copy-or-predict mechanism to generate drug sets. MoleRec [8] simulates interactions between drug substructures, aiming to identify specific substructures that significantly contribute to patient recovery. 4SDrug [10] proposes a symptom-based set-to-set framework aimed at recommending low-dose drug combinations to minimize DDI. VITA [7] investigates how past patient visits affect current ones, demonstrating that incorporating irrelevant past visits reduces drug recommendation performance. SHAPE [15] employs a visit-level set encoder with inducing points to deeply analyze the relationships and importance differences among medical codes within visits, effectively capturing visit-level representations. DrugDoctor [17] extracts information from multiple medical dimensions, enabling precise visit-level representations. LEADER [16] applies feature-level knowledge distillation to leverage the powerful semantic understanding capabilities of large language models in drug recommendation tasks, enhancing model performance while effectively addressing inference efficiency issues through distillation to a lightweight model. DNMDR [18] captures both temporal dynamics and structural dependencies through the constructed dynamic network to produce holistic patient representations.
Although significant progress has been made, existing models still directly input all historical visit records, ignoring the relationships between historical and current information, which introduces irrelevant data and reduces the model’s performance. Therefore, we propose the attribute-level history selection mechanism to effectively filter fine-grained historical information that is highly relevant to the patient’s current clinical condition.
Method
Problem definition
Electronic health record
The EHR for all patients can be represented as , where N is the number of patients. EHR for patient i can be denoted as a sequence of visits , where T(i) is the number of visits for the patient i. Each visit consists of multi-hot medical vectors (e.g., diagnoses, procedures, symptoms, and drugs). We use to denote the set of diagnoses, procedures, symptoms, and drugs, respectively. is the cardinality of the set. The t-th visit of patient i is defined as , where , , and . We describe the algorithm for a single patient and remove the superscript (i) when there is no ambiguity.
DDI graph
The symmetric binary adjacency matrices, denoted by , represent the DDI matrix. D stores information on known DDI from external knowledge. Here, = 1 means that there is an adverse reaction between the drug i and the drug j, so they should not be used together.
Drug recommendation task
The objective of the drug recommendation task is to generate a multi-hot prediction vector based on the patient’s historical visits and current visit information .
The main notations used in this paper are listed in Table 2.
Figure 3 shows the architecture of our model. which consists of three main components: (i) the patient representation module encodes the longitudinal health condition of patients; (ii) the drug representation module captures the detailed substructure information and overall molecular architecture of drugs; (iii) the predictor module makes a prediction.
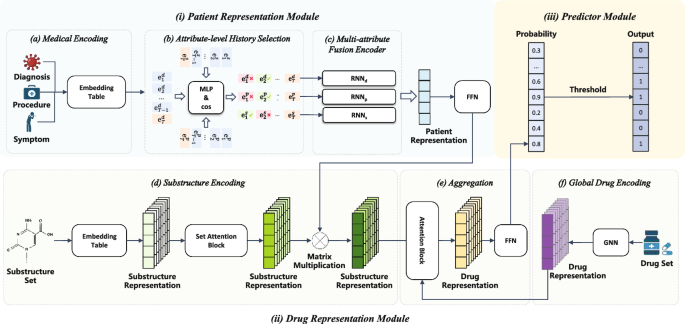
The EDRMM framework consists of: (i) Patient Representation Module, (ii) Drug Representation Module, (iii) Predictor Module
The patient representation module
We develop a comprehensive patient representation learning module that integrates detailed medical data from various visits and attributes. It can enhance drug recommendation tasks by leveraging multiple attributes and employing the attribute-level history selection mechanism for better modeling.
Medical encoding
In Sect. Problem definition, the t-th visit consists of medical multi-hot vectors of . To express the patient’s health status more comprehensively, we design three embedding tables: for diagnosis, for procedure, and for symptom, where dim is the dimension of medical embedding. We project the multi-hot medical vectors into the embedding space using vector-matrix multiplication.
Attribute-level history selection
We propose a novel attribute-level history selection mechanism for selecting relevant historical information. Different from visit-level selection in VITA [7], our model selects history visits from a more fine-grained perspective to retain more relevant information. We calculate the correlation score between each attribute of historical visits and the current attribute. In particular, considering a past visit of , and the current visit of , we calculate the correlation scores between and , and , and . As an example, we consider the calculation of the correlation score of between and .
Firstly, we employ a two-layer perceptron, concatenating and as input. The Multi-Layer Perceptron (MLP) learns an interactive representation based on these two features ( and ), ultimately generating the correlation coefficient score through the following formula:
where and are learnable weight matrices, while and are bias terms. represents the activation function, and [;] is the concatenation function.
Secondly, we utilize the cosine similarity algorithm to calculate the direction consistency of two vectors. It is independent of the vector lengths. Using cosine similarity to measure the correlation between two medical visits allows us to intuitively measure the similarity in the feature space.
where and are elements in the vectors and respectively, with a length of n.
We use the combined similarity calculation mechanism sim which combines MLP with cosine similarity to calculate the correlation score . The MLP captures complex nonlinear relationships and extracts deep features, while cosine similarity efficiently measures vector direction similarity. Finally, the final correlation coefficient score is obtained through the following formula:
where the function compresses correlation scores into the range [0,1]. Subsequently, we employ the same approach to calculate the correlation scores and for procedures and symptoms, respectively.
Based on the correlation score , we select relevant diagnoses from history visits using a parameter , forming a set of relevant diagnoses . If is greater than , it indicates a high correlation between and . Therefore, the definition of is as follows:
Similarly, we obtain and using the same approach (Using the same parameter). It is worth noting that, through attribute-level history selection mechanism, the sizes of , , and may vary.
Multi-attribute fusion encoder
To aggregate the relevant historical information , , and , along with current health states , , and , into the final patient representation, we use Recurrent Neural Network (RNN) [19] as a sequence encoder.
Medical data often suffers from missing data. For instance, certain medical records may solely contain diagnosis information, lacking procedure or symptom details, or only one of these types may be present. Using a single RNN to encode such data may compromise model performance when specific information is missing, thereby preventing the full utilization of available data. Inspired by [4], we propose a Triple-RNN aimed at learning patient representations from heterogeneous EHR data. Each RNN focuses on capturing the dependencies among medical codes for each attribute. This design ensures that all relevant information is adequately captured. Therefore, the model’s performance and generalization capability are enhanced. We input all patient information to RNN as follows:
The final representation r is achieved by concatenating the three hidden states:
The drug representation module
Some papers have indicated that the functional properties of drugs can be reflected through their substructures [20]. Consequently, previous models [8, 21] have explored this by decomposing drugs into substructures. Inspired by them, our model also leverages substructure information to model drugs. To represent drugs accurately, we harness the detailed information of their substructures and overall molecular architecture. In the following sections, we introduce the drug representation module that captures these essential elements, enabling robust drug encoding.
Global drug encoding
The molecular structure of each drug can be represented as a graph , where nodes V represent atoms and edges E represent chemical bonds. Recently, Graph Neural Networks (GNNs) have been widely applied to molecule-related tasks [5, 8, 21,22,23,24]. GNNs enable each node’s representation to incorporate not only its own information but also the structural and chemical properties of its neighboring nodes. Specifically, we utilize the L-layer Graph Isomorphism Network (GIN) [25] to obtain the global embedding of the drug. For each layer , each node receives messages from its neighboring nodes and updates its own representation:
where is a learnable parameter.
We obtain the global embedding g of the drug graph by applying a global mean pooling, which is defined as follows:
Substructure encoding
We employ the Breaking Retrosynthetically Interesting Chemical Substructures (BRICS) [26] mechanism to decompose all drugs in the set and generate a set of substructures . Similarly to drugs, substructures can also be represented as molecular graphs. To obtain the initial representation of substructures, we employ the GIN network to get the substructure embedding matrix . Drugs are typically composed of multiple substructures, which may exhibit various associations and interactions. Following [8], we utilize the Set Attention Block (SAB) [27] to update the representations of substructures, capturing the relationships and dependencies among them.
where is the substructure embeddings after applying the SAB.
In the context of a patient’s health condition, certain substructures play a significant role. So we utilize the patient’s final representation r as a query and employ a single-layer Feed-Forward Network (FFN) to compute the importance of each substructure for the patient. Then we update the substructure representation by multiplying the weights with the substructure embeddings. The formula is as follows:
where represents the learnable weight matrix, is the bias term, and is the updated substructure weight embeddings indicating the importance of each substructure to the patient.
Aggregation
For each drug k, we select its corresponding set of substructures . Since different substructures contribute differently to the drug’s properties, we use the global embedding of drug k as a query and apply an attention mechanism to compute the importance of each substructure within . The final representation of drug k, denoted as , is obtained by computing the weighted sum of the substructure embeddings:
where denotes the embedding of substructure s, and is the attention coefficient computed as:
where and are learnable weight matrices, and represents the embedding of substructure . Finally, we aggregate the representations of all drugs into a matrix .
The predictor module
We feed all drug representations into an FFN to obtain the prediction probabilities .
Here, represents the sigmoid function and represents the probability of each drug appearing in the prescription. A multi-hot prediction vector is then generated by choosing entries that exceed a predefined threshold .
Loss function
We perform optimization on all learnable parameters and utilize a combined loss function to achieve a more favorable balance between safety and accuracy during the training process. In this section, the combined loss will be introduced.
Multi-label prediction loss
In line with the prior works [4, 5, 9], the recommendation task is regarded as a multi-label classification task in our paper. Consequently, two prevalently utilized loss functions for multi-label classification, specifically, the binary cross-entropy loss and the multi-label marginal loss , are employed.
where the subscript t represents the t-th visit and the superscript (j) represents the j-th element of set.
DDI loss
For the purpose of controlling DDI within the context of combination drug use prediction. We define the DDI loss according to [4, 5]:
where gives the pairwise DDI probability.
We adopt the weight annealing algorithm proposed by [8], which dynamically adjusts the weight of DDI loss to balance safety and accuracy in drug recommendation.
where c is a current DDI rate and , are hyperparameters. The target DDI rate is denoted by . If , we optimize the DDI loss; otherwise, we optimize the multi-label loss without considering the DDI loss.
DDI risk regularization term
To better control the drug–drug DDI risk, we introduce a global DDI risk regularization term, denoted as . This term is designed to quantify the DDI risks in the predicted drug set by leveraging pre-existing global DDI knowledge. Specifically, it considers the normalized interaction probabilities between predicted drugs and computes a weighted summation over the drug pairs. By incorporating this term into the model, we aim to ensure that the generated recommendation minimizes DDI risks while maintaining accuracy. The risk formula is defined as follows:
The term is computed by summing the DDI risks across all predicted drug pairs (i, j) in the drug set . Here, the term represents the normalized interaction probability between drugs i and j, is the real DDI risk between these two drugs, and ensures that only upper-triangular drug pairs (i, j) are considered to avoid double-counting.
To compute the normalized interaction probability N, we first calculate the interaction probability matrix . This normalization ensures that the contribution of each drug to the DDI risk is proportional to its predicted probability, thus balancing the influence of frequent and infrequent drugs in the risk calculation.
Combined loss function
Given that drug interactions are intrinsically present in the real-world EHR data (e.g., the average DDI rate in the EHR dataset MIMIC-III is 0.0875), both accurate and inaccurate predictions might lead to an elevation in the DDI rate. During training, DDI rates typically increase alongside improvements in accuracy, yet our objective is to achieve an equilibrium between recommendation performance and safety. To address this, we propose a novel combined mechanism that integrates the weight annealing algorithm with a global DDI risk regularization term.
We combine loss function dynamically adjust weights at different stages of training to highlight the importance of multi-label prediction loss and DDI loss at various phases. Furthermore, DDI loss alone cannot fully leverage global drug interaction knowledge. The global risk regularization term incorporates pre-existing global drug interaction knowledge into the training process, enhancing the model’s ability to capture comprehensive global interaction information. We take the form of a weighted sum:
where , , are hyperparameters.
Overall, the training algorithm is detailed as Algorithm 1.

One training epoch of EDRMM
Dataset and settings
Dataset
MIMIC-IIIFootnote1 is a real-world EHR dataset [28], which comprises data from over 40,000 patients admitted to hospitals between 2001 and 2012, including demographics, procedures, drugs, laboratory results, etc. DDI relations are obtained from TWOSIDES [29] by following the data processing procedures in [11]. Table 3 summarizes the statistics of the processed dataset.
Baselines and metrics
Baselines contain the instance-based models of Logistic Regression (LR) and LEAP, and the longitudinal-based models of RETAIN, GAMENet, MICRON, SafeDrug, COGNet, MoleRec, VITA, SHAPE, LEADER and DrugDoctor. Following prior works [5, 6], we use four widely used metrics: Jaccard Similarity Score (Jaccard), F1 Score (F1), Precision Recall Area Under the Curve (PRAUC), and Drug–Drug-Interaction Rate (DDI).
Experimental settings
We divided the dataset into training, validation, and test sets as . We implement the model using Pytorch 2.0.1 and utilize the Adam optimizer with 50 epochs. For Triple-RNN, we use the gated recurrent unit (GRU) with 64 hidden units.
We obtained the optimal hyperparameters through grid search. Table 4 summarizes the range of hyperparameter search. The experimental results of dimension (dim), learning rate (lr), dropout rate (dp), , and are shown in Fig. 4, respectively. Specifically, we selected the hyperparameters based on the performance on the validation set, with the following values: lr = 0.0005, dp = 0.8, = 2.5, = 0.5, and dim = 64. We set the number of GIN layers L to 4. Additionally, following the standard practice for binary classification, we set the threshold = 0.5, = 0.5. The same as MoleRec, we set the weight = 0.95, and the target DDI rate = 0.06.
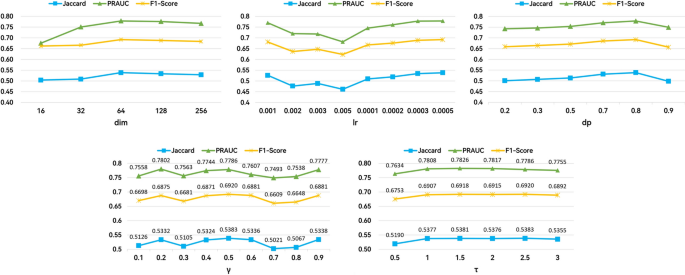
The experimental results of hyperparameters
We conducted our experiments on a Linux server equipped with an NVIDIA GeForce RTX 3090 GPU. All baselines in our paper were implemented using the optimized parameters as described in the respective references.
Results
Table 5 shows the experimental results on the MIMIC-III dataset, we have the following observations:
- (1)Although DrugDoctor shows slightly better performance in PRAUC, our model demonstrates superior results in all other metrics. This is largely because DrugDoctor uses all historical visit records in the patient representation module without considering their relevance to the current visit, while EDRMM leverages attribute-level historical information that is more relevant to the patient’s current clinical condition, thus avoiding interference from irrelevant historical data. Additionally, while DrugDoctor constructs patient representations using only diagnosis and procedure information, our EDRMM enhances these representations by incorporating additional EHR attributes, which further improves the accuracy of the recommendations.
- (2)EDRMM demonstrates significant improvements over VITA. This advantage is primarily attributed to our model’s attribute-level history selection mechanism, which enables more precise retention of valuable information. VITA’s relevant-visit selection mechanism filters out entire visits if they are deemed irrelevant. Additionally, VITA relies on the overall representation of drugs, overlooking substructures that more effectively reflect their functional properties.
- (3)The traditional LR model predicts fewer drugs on average but yields a higher DDI rate, suggesting that the recommended drugs exhibit greater interactions, potentially indicating higher risks. LEAP demonstrates the lowest accuracy among all models. Both LR and LEAP are instance-based models that consider only the current visit and do not utilize patient historical information, making them less effective for drug recommendation compared to longitudinal-based models that leverage such information.
- (4)We further investigate the impact of the number of visits on the performance of EDRMM. Given that most patients in the MIMIC-III visit the hospital fewer than five times (Fig. 5), we analyze the first five visits for each patient in the preprocessed dataset and the test set, by following an existing model [9]. We compared our model with two strong baselines: MoleRec and DrugDoctor. Figure 6 shows that EDRMM demonstrates remarkable robustness across varying numbers of visits. When there is no patient history (only a single visit record), EDRMM consistently outperforms MoleRec and DrugDoctor. All models demonstrate improved performance for patients with a second visit, which may be attributed to the relatively high proportion of such patients in the MIMIC-III. Although DrugDoctor performs relatively well for patients with a second or third visit, its performance is unstable, exhibiting significant fluctuations as the number of visits increases.
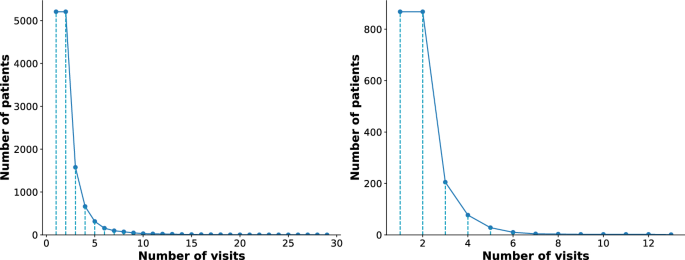
The histogram of hospital visits of patients in the preprocessed MIMIC-III dataset (left) and test dataset (right)
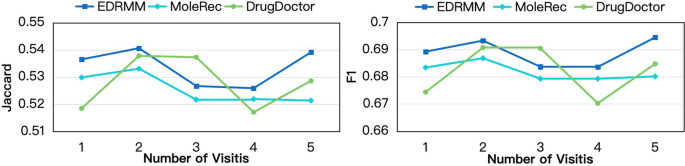
EDRMM is compared with DrugDoctor and MoleRec in terms of Jaccard and F1 across different numbers of visits
Discussion
Ablation study
We conduct an ablation study to evaluate the three core innovations of our model. The details of the ablation settings are as follows:
- Multi-attribute Modeling. We implement three variants to examine the contribution of different attributes:
- : remove diagnosis from the patient representation while retaining procedure and symptom attributes.
- : remove procedure from the patient representation while retaining diagnosis and symptom attributes.
- : remove symptom from the patient representation while retaining diagnosis and procedure attributes.
- Attribute-level History Selection Mechanism. The w/o sim variant removes the attribute-level history selection mechanism to assess its role in filtering relevant historical information.
- DDI Risk Regularization Term. The variant removes the DDI risk regularization term and uses the weighted annealing algorithm, evaluating the benefit of our adaptive DDI regularization.
This structured experimental design, summarized in Table 6, provides clear evidence for each component’s contribution to the overall model performance.
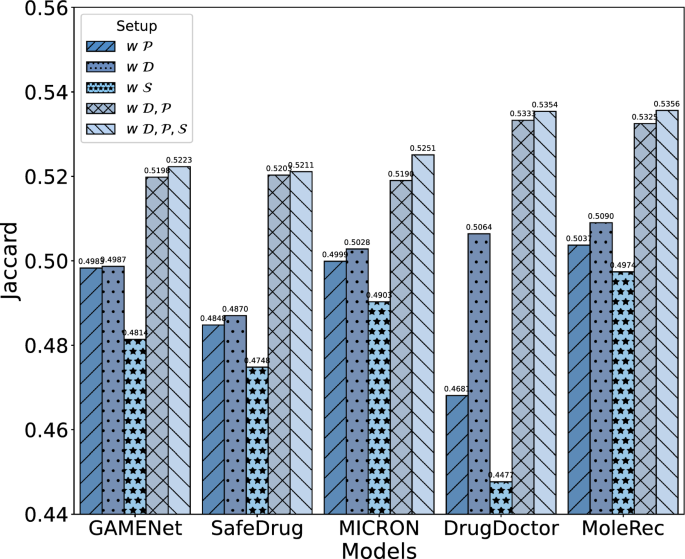
Ablation study for the impact of EHR attributes on patient representation in items of Jaccard
Effectiveness of multi-attribute modeling
To evaluate whether a multi-attribute approach improves drug recommendation and to identify which attributes are most beneficial, we conducted a series of experiments on four baseline models: GAMENet, SafeDrug, MICRON, DrugDoctor and MoleRec. The following observations address the two sub-questions of Q1.
The experimental results in Fig. 7 show that incorporating multiple attributes consistently enhances drug recommendation performance across all models. Notably, when only one attribute (e.g., diagnosis, procedure, or symptom) is used individually, the models still maintain a certain level of accuracy, indicating that each attribute contributes positively to the overall performance. This confirms that the incorporation of multi-attribute is effective. The results further indicate that adding more types of EHR attributes improves drug recommendation performance. For example, when the symptom attribute is added to models using only diagnosis and procedure attributes ( vs. ), performance is moderately enhanced, as shown in Fig. 7. This demonstrates that incorporating more attributes, when relevant, leads to incremental improvements in model performance.
As seen in Fig. 7 and Table 6, excluding diagnosis information leads to the most significant decline in performance across all models. This finding highlights the vital role of diagnosis attributes in constructing effective patient representations. Procedure attributes contribute by providing supplementary details on treatments, while symptom attributes capture patient-reported conditions, both positively enhancing performance, though their impact is less pronounced compared to diagnosis.
Effectiveness of the attribute-level history selection mechanism
We remove the attribute-level history selection mechanism and instead utilize all historical information as input. In Table 6, the recommendation performance declines significantly after removing the attribute-level history selection mechanism. This mechanism enables the model to select the most relevant information from the historical visit records of the current patient, thereby improving the accuracy of the recommendation.
Effectiveness of the DDI risk regularization term
In Table 6, removing the regularization term from our model results in decline across all model metrics, and causes a noticeable increase in the DDI rate. This demonstrates that the global DDI risk regularization term is critical for balancing the accuracy and safety of drug recommendation.
Efficiency
We evaluate the efficiency of each model in terms of parameter size, training time, and GPU memory usage. As reported in Table 7, all experiments are conducted under the same hardware configuration for consistency. The training time refers to the average duration per epoch, and the GPU memory usage reflects the allocated memory during training.
EDRMM achieves a favorable balance between performance and computational cost. With only 0.59M parameters and 26.06MB of allocated GPU memory during training, it remains lightweight compared to resource-heavy models such as SHAPE. The model also exhibits efficient training behavior, completing each epoch within a moderate runtime. These results highlight the practicality of EDRMM in scenarios with limited computational resources.
Case study
To visually demonstrate the advantages of EDRMM, we randomly sample cases and analyze their drug prediction results relative to ground-truth prescriptions. Due to space constraints, diagnosis and procedure data are abbreviated using International Classification of Diseases (ICD) codes [30], while drug data are abbreviated using Anatomical Therapeutic Chemical (ATC) codes [31].
We randomly selected a patient from the MIMIC-III dataset, referred to as patient A. We analyzed the number of drugs recommended by three top-performing models (including MoleRec, DrugDoctor, and EDRMM) for patient A and their DDI rates across different visits. As shown in Fig. 8, the line charts display the DDI rates of the three models across patient A’s visits. The results indicate that EDRMM not only recommends the most drugs for the doctor but also maintains the lowest DDI rate. Across the four visits, EDRMM recommended a total of 92 drugs, with only 18 drug combinations exhibiting interactions.
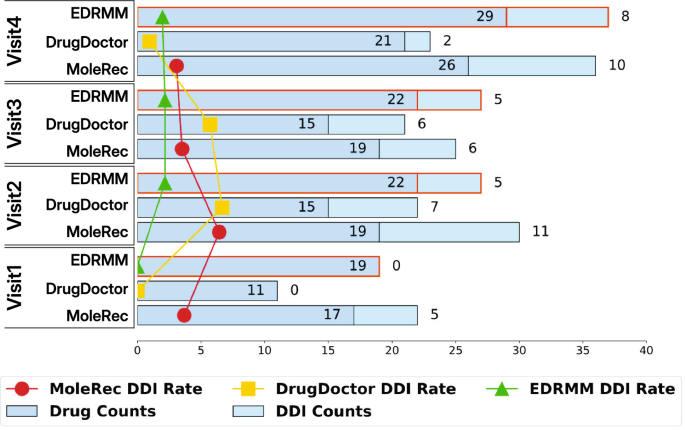
An example of the number of recommended drugs, DDI counts, and DDI rates for patient A
In addition, Table 8 shows the recommendation results of the three models for patient A’s first and second visits. The results indicate that EDRMM demonstrates better recommendation effectiveness and achieves a higher Jaccard similarity.
To further verify the functionality of our attribute-level history selection mechanism in more complex clinical scenarios, we randomly select a patient (referred to as patient B) from the test set, who underwent four hospital visits. Using the attribute-level history selection mechanism, we illustrate the model’s choices in retaining or discarding information from each visit, as shown in Table 9. The second and third visits provide more comprehensive details, reflecting the patient’s condition progression and treatment process. Given the high degree of similarity with the third visit, our attribute-level history selection mechanism retained all information for this visit. The specific reasons are as follows:
- (1)Diagnosis attribute The current visit and the third visit share many of the same diagnoses. For example, both visits include 0389 (Septicemia NOS), 2720 (Pure hypercholesterolemia), 42731 (Atrial fibrillation), 496 (Chronic airway obstruction NEC), and 56981 (Intestinal fistula).
- (2)Procedure attribute Both visits involve complex medical interventions. During the third visit, procedures included 8847 (Contrast abdominal arteriography NEC) and 3979 (Other endovascular procedures on other vessels), while during the current visit, procedures encompass 3893 (Venous catheterization NEC), 966 (Enteral infusion of nutritive substances), 3891 (Arterial catheterization), 3404 (Insertion of the intercostal catheter) and 3491 (Thoracentesis). Although the procedures differ, they both necessitate specialized medical personnel for execution. This suggests that both visits may involve relatively severe or intricate health issues, with similarity lying in the complexity and intensity of medical interventions rather than specific procedure codes.
- (3)Symptom attribute The symptoms observed during the current visit, such as fever, pain, and dizziness, also appear on the third visit. Therefore, the symptoms from the third visit are highly similar to the current symptoms and hold significant reference value.
Limitations
There are still some limitations in this study. First, although we incorporated multi-granularity and multi-attribute medical auxiliary information, numerous clinically valuable data modalities (e.g., vital signs and laboratory test records) remain underexplored. Second,adopting more advanced graph neural networks (e.g., graph Transformers) for molecular data processing could potentially yield more effective drug representations, thus improving recommendation performance. Future work will focus on exploring more clinical data modalities and adopting advanced graph neural network techniques to improve drug recommendation performance.
Conclusions
We propose EDRMM, a state-of-the-art model addressing key challenges in drug recommendation. Specifically, EDRMM tackles the limitations of ignoring the relationship between historical and current visits and the underutilization of EHR information. To capture the relationship between historical and current visits, we propose an attribute-level history selection mechanism that retrieves relevant historical information while filtering out noise. By integrating multi-attribute data and employing separate encoding strategies, the model fully leverages EHRs to create comprehensive patient representations. The hybrid loss function, which includes a global DDI risk regularization term, achieves a balance between accuracy and safety in drug recommendation. Experimental results on the MIMIC-III dataset validate the effectiveness and superiority of EDRMM.
Availability of data and materials
The datasets analysed during the current study are available in the MIMIC-III Clinical Database repository, https://physionet.org/content/mimiciii/1.4/.[28] The source code is available at GitHub: https://github.com/ricartojason/EDRMM/tree/main.
Notes
-
https://physionet.org/content/mimiciii/1.4/.
Abbreviations
- EHR:
- Electronic Health Record
- DDI:
- Drug–Drug Interaction
- MLP:
- Multi-Layer Perceptron
- RNN:
- Recurrent Neural Network
- GNN:
- Graph Neural Network
- GIN:
- Graph Isomorphism Network
- BRICS:
- Breaking Retrosynthetically Interesting Chemical Substructures
- SAB:
- Set Attention Block
- FFN:
- Feed-Forward Network
- LR:
- Logistic Regression
- GRU:
- Gated Recurrent Unit
- ICD:
- International Classification of Disease
- ATC:
- Anatomical Therapeutic Chemical
References
-
Lee CS, Lee AY. Clinical applications of continual learning machine learning. Lancet Digital Health. 2020;2(6):e279–81. https://doi.org/10.1016/s2589-7500(20)30102-3.
-
Obermeyer Z, Emanuel EJ. Predicting the future-big data, machine learning, and clinical medicine. N Engl J Med. 2016;375(13):1216. https://doi.org/10.1056/nejmp1606181.
-
Shamout F, Zhu T, Clifton DA. Machine learning for clinical outcome prediction. IEEE Rev Biomed Eng. 2020;14:116–26. https://doi.org/10.1109/RBME.2020.3007816.
-
Shang J, Xiao C, Ma T, Li H, Sun J. Gamenet: Graph augmented memory networks for recommending medication combination. In: Proceedings of the AAAI conference on artificial intelligence. 2019;33:1126–33.
-
Yang C, Xiao C, Ma F, Glass L, Sun J. SafeDrug: Dual Molecular Graph Encoders for Safe Drug Recommendations. In: Proceedings of the thirtieth international joint conference on artificial intelligence, IJCAI 2021;2021.
-
Yang C, Xiao C, Glass L, Sun J. Change Matters: Medication Change Prediction with Recurrent Residual Networks. In: Proceedings of the thirtieth international joint conference on artificial intelligence, IJCAI 2021;2021.
-
Kim T, Heo J, Kim H, Shin K, Kim SW. VITA: ‘Carefully Chosen and Weighted Less’ Is Better in Medication Recommendation. In: Proceedings of the AAAI conference on artificial intelligence. 2024;38:8600–7. Available from: https://doi.org/10.1609/aaai.v38i8.28704.
-
Yang N, Zeng K, Wu Q, Yan J. MoleRec: combinatorial drug recommendation with substructure-aware molecular representation learning. In: Proceedings of the ACM web conference. 2023;2023:4075–85.
-
Wu R, Qiu Z, Jiang J, Qi G, Wu X. Conditional generation net for medication recommendation. In: Proceedings of the ACM web conference. 2022;2022:935–45.
-
Tan Y, Kong C, Yu L, Li P, Chen C, Zheng X, et al. 4sdrug: Symptom-based set-to-set small and safe drug recommendation. In: Proceedings of the 28th ACM SIGKDD conference on knowledge discovery and data mining; 2022;3970–3980.
-
Sun H, Xie S, Li S, Chen Y, Wen JR, Yan R. Debiased. Longitudinal and Coordinated Drug Recommendation through Multi-Visit Clinic Records. In: Advances in Neural Information Processing Systems; 2022.
-
Gong F, Wang M, Wang H, Wang S, Liu M. SMR: medical knowledge graph embedding for safe medicine recommendation. Big Data Res. 2021;23: 100174. https://doi.org/10.1016/j.bdr.2020.100174.
-
Zhang Y, Chen R, Tang J, Stewart WF, Sun J. LEAP: learning to prescribe effective and safe treatment combinations for multimorbidity. In: Proceedings of the 23rd ACM SIGKDD international conference on knowledge Discovery and data Mining; 2017;1315–1324.
-
Choi E, Bahadori MT, Sun J, Kulas J, Schuetz A, Stewart W. Retain: an interpretable predictive model for healthcare using reverse time attention mechanism. Adv Neural Inf Process Syst. 2016;29.
-
Liu S, Wang X, Du J, Hou Y, Zhao X, Xu H, et al. SHAPE: a sample-adaptive hierarchical prediction network for medication recommendation. IEEE J Biomed Health Inform. 2023;27(12):6018–28. https://doi.org/10.1109/JBHI.2023.3320139.
-
Liu Q, Wu X, Zhao X, Zhu Y, Zhang Z, Tian F, et al. Large language model distilling medication recommendation model. arXiv preprint arXiv:2402.02803. 2024;.
-
Kuang Y, Xie M. DrugDoctor: enhancing drug recommendation in cold-start scenario via visit-level representation learning and training. Brief Bioinform. 2024;25(6):bbae464. https://doi.org/10.1093/bib/bbae464.
-
Liu G, Yu X, Liu Z, Li X, Fan X, Zheng X. DNMDR: dynamic networks and multi-view drug representations for safe medication recommendation. Available from: https://arxiv.org/abs/2501.08572.
-
Schuster M, Paliwal KK. Bidirectional recurrent neural networks. IEEE Trans Signal Process. 1997;45(11):2673–81. https://doi.org/10.1109/78.650093.
-
Nyamabo AK, Yu H, Shi JY. SSI-DDI: substructure-substructure interactions for drug-drug interaction prediction. Brief Bioinform. 2021. https://doi.org/10.1093/bib/bbab133.
-
Wu J, Qian B, Li Y, Gao Z, Ju M, Yang Y, et al. Leveraging multiple types of domain knowledge for safe and effective drug recommendation. In: Proceedings of the 31st ACM international conference on information & knowledge management. CIKM ’22. New York, NY, USA: Association for Computing Machinery; 2022. p. 2169-2178.
-
Rong Y, Bian Y, Xu T, Xie W, Wei Y, Huang W, et al. Self-supervised graph transformer on large-scale molecular data. Adv Neural Inf Process Syst. 2020;33:12559–71.
-
Hao Z, Lu C, Huang Z, Wang H, Hu Z, Liu Q, et al. ASGN: An active semi-supervised graph neural network for molecular property prediction. In: Proceedings of the 26th ACM SIGKDD international conference on knowledge discovery & data mining; 2020;731–752.
-
Yu Z, Gao H. Molecular representation learning via heterogeneous motif graph neural networks. In: International Conference on Machine Learning. PMLR; 2022;25581–25594.
-
Xu K, Hu W, Leskovec J, Jegelka S. How powerful are graph neural networks? In International Conference on Learning Representations. 2018;.
-
Degen J, Wegscheid-Gerlach C, Zaliani A, Rarey M. On the Art of Compiling and Using “Drug-Like” Chemical Fragment Spaces. ChemMedChem. 2008;1503-1507. https://doi.org/10.1002/cmdc.200800178.
-
Lee J, Lee Y, Kim J, Kosiorek A, Choi S, Teh YW. Set transformer: A framework for attention-based permutation-invariant neural networks. In: International conference on machine learning. PMLR; 2019;3744–3753.
-
Johnson AE, Pollard TJ, Shen L, Lehman LWH, Feng M, Ghassemi M, et al. MIMIC-III, a freely accessible critical care database. Sci Data. 2016;3(1):1–9. https://doi.org/10.1038/sdata.2016.35.
-
Tatonetti NP, Ye PP, Daneshjou R, Altman RB. Data-Driven Prediction of Drug Effects and Interactions. Sci Transl Med. 2012. https://doi.org/10.1021/acs.oprd.9b00202.s001.
-
Slee VN.: The international classification of diseases: ninth revision (ICD-9). American College of Physicians.
-
Cheng X, Zhao SG, Xiao X, Chou KC. iATC-mHyb: a hybrid multi-label classifier for predicting the classification of anatomical therapeutic chemicals. Oncotarget. 2017;8(35):58494.
